ORA™ SEE qPCR Green ROX H Mix
Applications
- Dye-based qPCR.
- qPCR from gDNA, cDNA, viral DNA, low copy number genes.
- Relative gene expression analysis, absolute quantification.
Benefits
- Universal - both standard and fast cycling.
- Excellent for GC or AT rich templates.
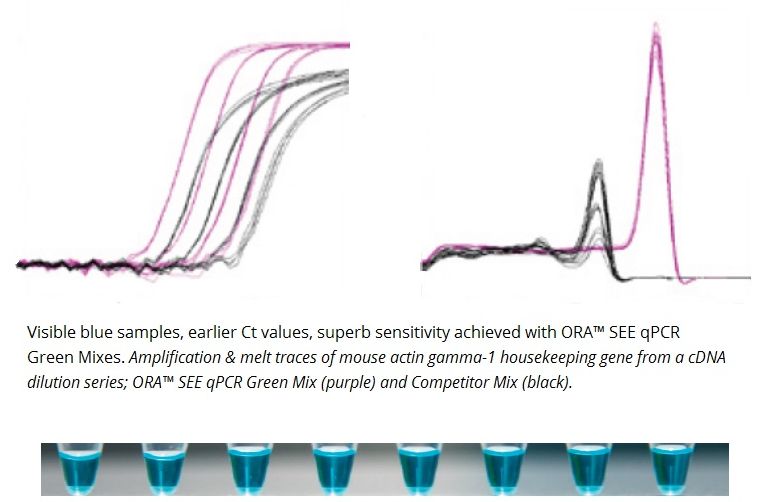
- Highest sensitivity, rapid extension, early Ct values.
- Inert blue dye for a better sample visibility and tracking.
ORA™ SEE qPCR Green ROX H Mix







- Description highQu qPCR master mixes are based on the small molecular inhibitor technology Hot Start PCR allowing to achieve highest sen… More
- Protocols Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X Download MSDS ORA™ SEE qPCR Green RO More
- Specifications Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X Download MSDS ORA™ SEE qPCR Green RO More
- Resources Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X Download MSDS ORA™ SEE qPCR Green RO More
- Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X
- Download MSDS ORA™ SEE qPCR Green ROX H Mix, 2X
- Need a lot-specific Certificate of Analysis? E-mail us at info@highQu.com
- Want custom formulations or bulk sizes? E-mail us at info@highQu.com, and check our OEM offers
- Have more specific questions? Contact us
- Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X
- Download MSDS ORA™ SEE qPCR Green ROX H Mix, 2X
- Need a lot-specific Certificate of Analysis? E-mail us at info@highQu.com
- Want custom formulations or bulk sizes? E-mail us at info@highQu.com, and check our OEM offers
- Have more specific questions? Contact us
- Download Protocol and Specifications - Product Insert ORA™ SEE qPCR Green ROX H Mix, 2X
- Download MSDS ORA™ SEE qPCR Green ROX H Mix, 2X
- Download highQu Catalogue of Premium Research Tools
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Product Pricelist (DE 2021)
- Download all highQu Product Inserts and MSDS sheets
- Have questions? Contact us
Related Products