SampleIN™ Direct PCR Kit
Applications
- Direct PCR without template purification.
- Mouse genotyping and knockout analysis.
- Direct PCR from blood, mouse tail or ear, mammalian tissues (including FFPE).
- Direct PCR from hair follicle, buccal swabs, plant samples.
Benefits
- Ready-to load PCR in 50 minutes without template purification.
- Single-tube 15 min DNA extraction combined with fast hot-start PCR.
- Red dye in the PCR master mix for direct gel loading.
- High yields under standard or fast cycling conditions.
- Success with GC/AT rich templates.
SampleIN™ Direct PCR Kit











- Description SampleIN™ Direct PCR Kit is a premium tool for a fast direct PCR eliminating the need of tedious template purification. The… More
- Protocols Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit Ask for a Sample today Have technical questio More
- Specifications Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit Download MSDS SampleIN™ Direct PCR Kit Need a More
- Resources Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit Download MSDS SampleIN™ Direct PCR Kit Downlo More
Product information "SampleIN™ Direct PCR Kit"
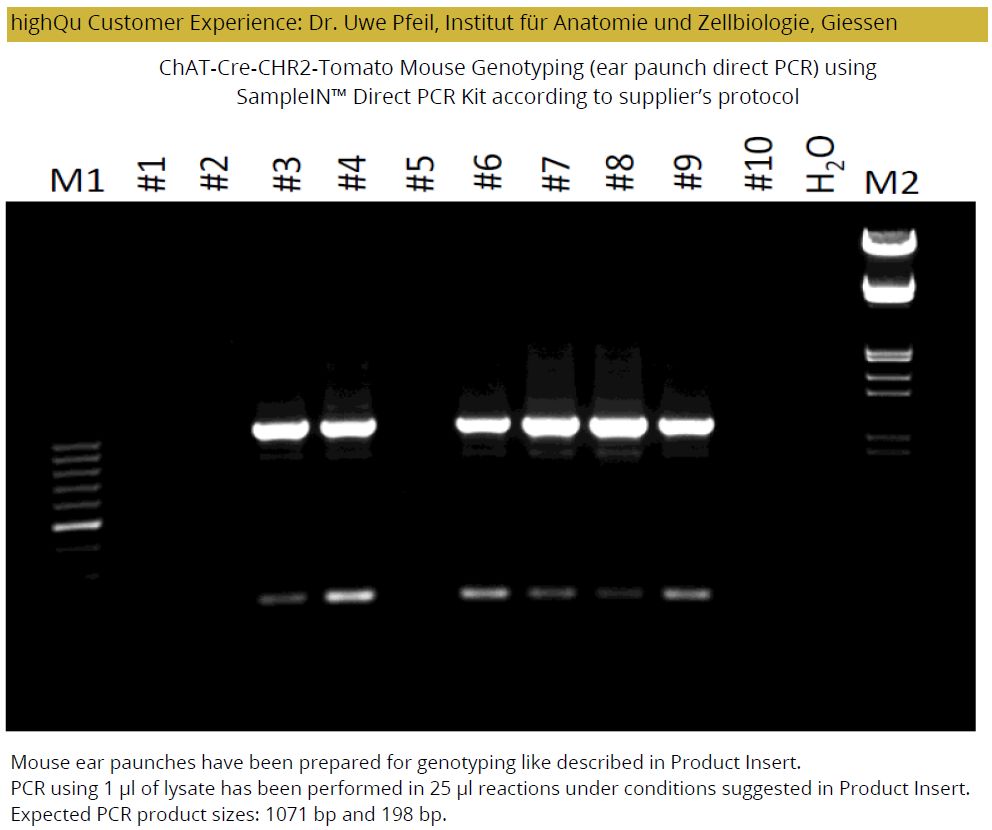
SampleIN™ Direct PCR Kit is a premium tool for a fast direct PCR eliminating the need of tedious template purification. The kit is excellent for direct PCR from mouse tail or ear, mammalian tissues, hair follicle, buccal swabs and blood.
Rapid 15 min DNA extraction using DPK Lysis and Protease Buffers in a single tube generates PCR template extract which is further amplified under fast cycling conditions with a hot-start Taq master mix that includes red dye for direct gel loading.
In a 2% agarose TAE gel the red dye migrates with~350 bp DNA, in 1% agarose TAE gel with ~ 600 bp DNA fragments.
The ALLin™ HS Red Taq Mastermix includes a hot start Taq DNA Polymerase what ensures high yield, specific, low background amplification. Mix components allow for a fast PCR cycling and increase success when working with complex templates or multiplexing. Generated A-tailed PCR products are suitable for ligating into TA cloning vectors, sequencing and other applications.
- Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit
- Ask for a Sample today
- Have technical questions? Contact us
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit
- Download MSDS SampleIN™ Direct PCR Kit
- Need a lot-specific Certificate of Analysis? E-mail us at info@highQu.com
- Want custom formulations or bulk sizes? E-mail us at info@highQu.com, and check our OEM offers
- Have more specific questions? Contact us
- Download Protocol and Specifications - Product Insert SampleIN™ Direct PCR Kit
- Download MSDS SampleIN™ Direct PCR Kit
- Download highQu Catalogue of Premium Research Tools
- Download a list of Publications mentioning highQu products
- Or see how others use our products at bioz.com or google scholar
- Download Product Pricelist (DE 2021)
- Download all highQu Product Inserts and MSDS sheets
- Have questions? Contact us
Related Products
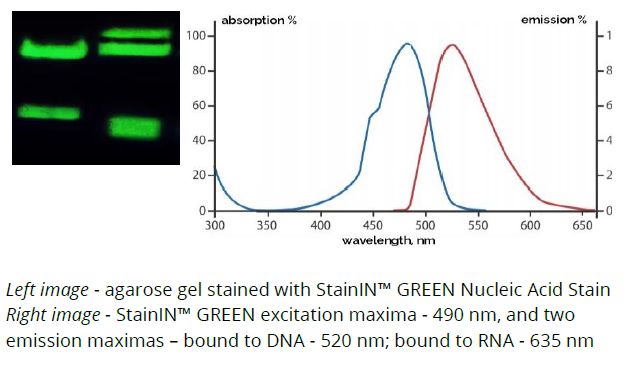
StainIN™ GREEN Nucleic Acid Stain
€98.00*
StainIN™ GREEN Nucleic Acid Stain is a significantly safer alternative to ethidium bromide.
It is same easy to use, 4X as sensitive (under Blue LED) and much more secure. More economical as other green dyes, this stain can be also used and disposed with less environmental and health concerns compared to ethidium bromide.
It is a fluorescent dye that allows detection of >0,1 ng of DNA in both agarose and polyacrylamide gels. It binds to both dsDNA, ssDNA and RNA and emits green fluorescence when bound to DNA and red fluorescence when bound to RNA detectable under the UV or Blue light and documented with same filters as similar green dyes.
StainIN™ GREEN is ideal for DNA extraction from gels for cloning. Smaller than ethidium bromide carcinogenicity has been proved by Ames-test. Mammalian cell mutagenicity tests, both mouse marrow erythrocyte micronucleus and spermatocyte chromosomal aberration tests gave negative mutagenicity results.
Carcinogenicity and toxicity of the widely used Ethidium Bromide (EtBr) is one of the biggest safety and environmental problems in the labs where gel electrophoresis is a daily routine. The less toxic nucleic acid dyes address this problem and minimize concerns.
The StainIN™ GREEN (SYBR Green-like stains alternative), (SYBR® Green is a registered trade mark of Molecular Probes Inc.) and StainIN™ eco-RED Nucleic Acid Stains by highQu provide a much less-carcinogenic, significantly safer alternative to EtBr while delivering even several times higher detection sensitivity. Same like EtBr, these stains are used during the process of electrophoresis in an agarose gel or PAAG and in the electrophoresis running buffer.
Nucleic acid stains are typically divided into two groups – they are either intercalating dyes or minor groove binders. The StainIN™ GREEN and StainIN™ eco-RED Nucleic Acid Stains belong to the group of intercalating dyes that bind to NA and produce the fluorescence signal. Same like many other suppliers of similar products, highQu retains the right to keep the precise chemical information undisclosed and proprietary.
The StainIN™ GREEN Nucleic Acid Stain as well as the StainIN™ eco-RED Nucleic Acid Stain are both supplied in water. Detailed safety information is available in MSDS sheets of the stains.
Despite proven less mutagenicity, all nucleic acid stains, including SYBR® Green or other shall be handled with care.
Used electrophoresis buffer with a stain as well as melted gels shall be run through approved (carbon) filters. If the absence of residual fluorescence is confirmed, the liquids can be disposed with plenty of water down the drain.
Alternatively, all used liquids can be autoclaved as the StainIN™ GREEN and StainIN™ eco-RED Nucleic Acid Stains degrade during the long exposure to above 100ºC temperature.
The safety office shall be consulted periodically to match local regulations, as they vary and change.
Variants from €108.00*
ALLin™ HS Red Taq Mastermix
€189.50*
highQu ALLin™ Hot Start Taq DNA Polymerase is the superior sensitive enzyme. The activity at room temperature is blocked by small molecular inhibitor. Enzyme becomes active only after heating what allows for highly specific and extremely sensitive amplification, no primer dimer formation and no background.
In combination with the optimized ALLin™ buffer enzyme provides higher success rates in demanding PCR applications like amplification of crude, complex or longer templates and fast cycling.
ALLin™ Hot Start Taq DNA Polymerase has the same PCR accuracy like Taq DNA Polymerase, 4.5 x 10^4 (nucleotides incorporated before the error occurs) and produces A-tailed products suitable for ligating into TA cloning vectors.
The convenience of ALLin™ Hot Start Taq DNA Polymerase is maximized by the use of 2X Red colored Mastermix providing the additional advantage of reduced pipetting and minimized errors. The mastermix is even supplied with PCR Water, and the only thing to add is the template with primers.
ALLin™ HS Red Taq Mastermix, 2X is premixed with red dye and density reagents for direct loading on the gels after the PCR.
In a 2% agarose TAE gel the dye migrates with~350 bp DNA, in 1% agarose TAE gel with ~ 600 bp DNA fragments.
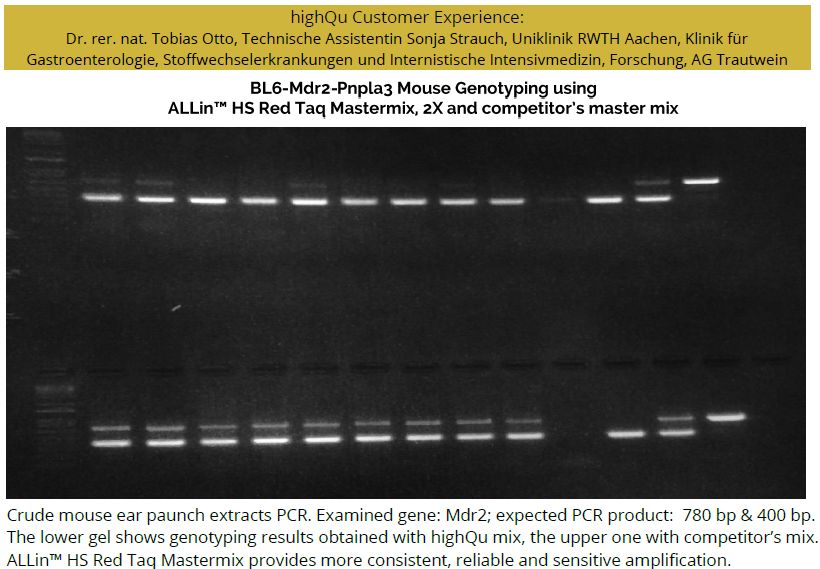
ALLin™ HS Red Taq Mastermix, 2X is also a key component in highQu SampleIN™ Direct PCR Kit (DPK0101/5), ensuring outstanding PCR results with crude samples.
Variants from €208.50*
Take5™ HR DNA Ladder
€109.50*
highQu ready to use DNA ladders are mixtures of highly purified plasmid digests and PCR products. The high ladder purity allows for exceptional stability and room temperature storage. Ladders are ready to be directly loaded on agarose gels and are supplied with the loading dye solution for probe DNA. They provide sharp bands for DNA sizing and allow for approximate DNA quantification by comparing the amount of DNA of the band similar in size to your sample band.
Variants from €120.50*
100 mM dNTP Set
€129.50*
highQu dNTP sets and mixes meet all highest industry standards and allow for unrivaled performance of your PCR and other DNA synthesis, labeling or sequencing reactions.
Produced under the stringent quality monitoring conditions, they guaranty reproducible results. More than 99% HPLC purity eliminates inhibitions of PCR and allows for increased yields with higher dNTP concentrations.
Exceptional stability of dNTPs allows for short term ambient temperature shipments, short term room temperature storage or long PCR of more than 30 kb targets, as well as long amplifications exceeding 40 cycles.
Variants from €142.50*